LandSCENT (Landscape Single Cell Entropy)
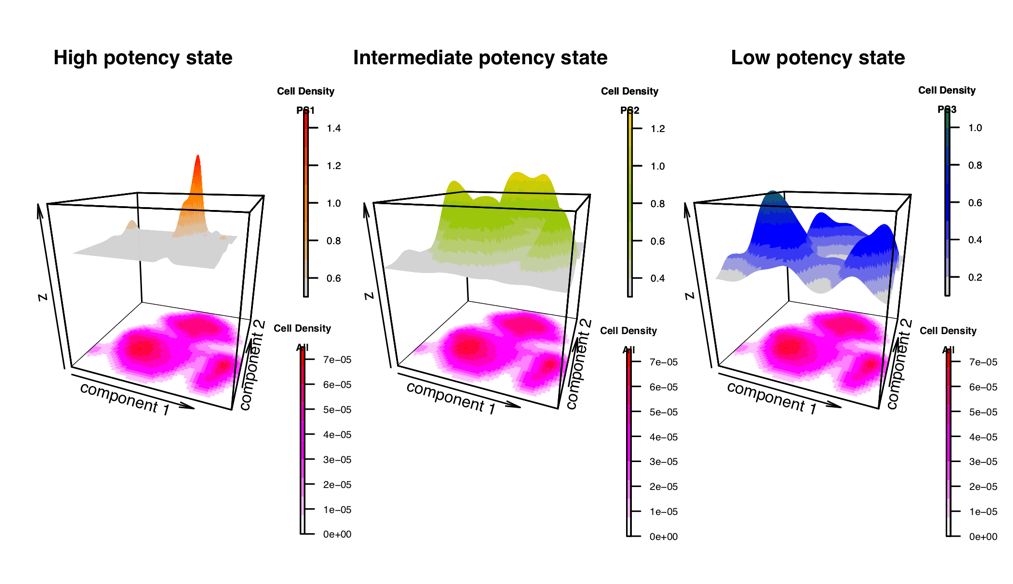
LandSCENT is an R-package for the analysis of single-cell RNA-Seq data. One main purpose of this package is to provide a means of estimating the differentiation potency of single cells without the need to assume prior biological knowledge (e.g. marker expression or timepoint). As such, it may provide a more unbiased means for assessing potency or pseudotime. LandSCENT also integrates cell density with potency distribution to dissect cell types across all potency states and generates high-quality figures to show it.
Maintainer: Weiyan Chen
Citation
Chen, W., Morabito, S.J., Kessenbrock, K. et al. "Single-cell landscape in mammary epithelium reveals bipotent-like cells associated with breast cancer risk and outcome." Communications Biology 2, 306 (2019).
Teschendorff, Andrew E, and Tariq Enver. 2017. "Single-cell entropy for accurate estimation of differentiation potency from a cell's transcriptome." Nature Communications 8 (1): 15599.
Teschendorff, Andrew E, Peter Sollich, and Reimer Kuehn. 2014. "Signalling entropy: A novel network-theoretical framework for systems analysis and interpretation of functional omic data." Methods 67 (3): 282.
Chen, Weiyan and Andrew E. Teschendorff. "Estimating Differentiation Potency of Single Cells Using Single-Cell Entropy (SCENT)." Computational Methods for Single-Cell Data Analysis. Humana Press, New York, NY, 2019. 125-139.
Shi, Jifan, Andrew E. Teschendorff, Weiyan Chen, Luonan Chen, and Tiejun Li. "Quantifying Waddington's epigenetic landscape: a comparison of single-cell potency measures." Briefings in bioinformatics (2018).
Installation
To install this package, start R (version "3.6") and enter:
if (!requireNamespace("devtools", quietly = TRUE))
install.packages("devtools")
devtools::install_github("ChenWeiyan/LandSCENT")
Documentation
HTML: LandSCENT Vignette